AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Paraview label format11/27/2023 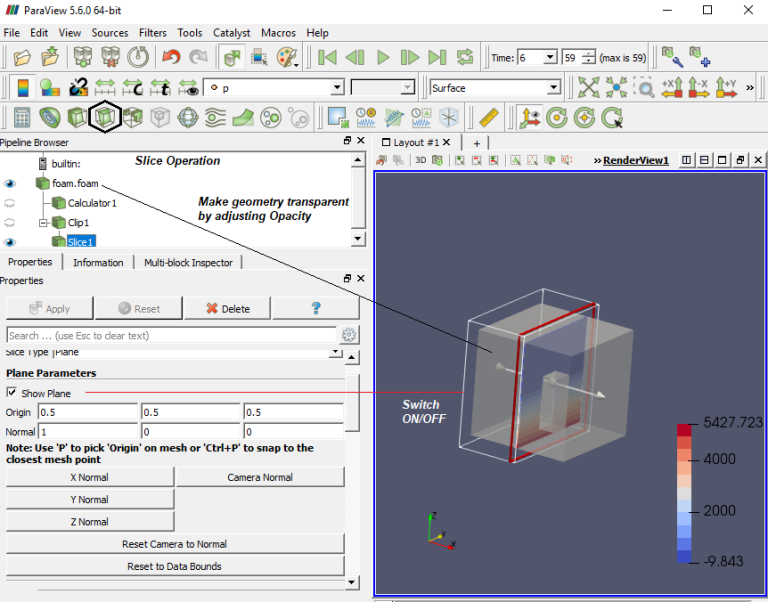
Additionally one can change the length of the legend by dragging the end-markers shown when one hovers the mouse pointer over the legend. One can click and drag the legend to place it interactively at any position in the view. One can control the characteristics of the color legend using the Edit color legend properties button. On one side are the automatically generated labels, while on the other side are the annotations, including the start and end annotation which corresponding to the data range being mapped. By default, the title is typically the name of the array (and component number or magnitude for non-scalar data arrays) being mapped. This button affects the visibility of the legend in the active view.įigure 2 shows the various components of the color legend. One can toggle the visibility of the color legend corresponding to the transfer function being shown/edit in the Color Map Editor by using the Show/Hide color legend button near the top of the panel. In this post, we take a look at color legend in 3D views and controlling the labels shown on the legend using annotations.įigure 1: Color Map Editor panel showing buttons that affect the color legend The Color Legendįigure 2: Color LegendThe color legend, also known as scalar bar or color bar, is designed to provide the user information about which color corresponds to what data value in the rendered view. CellML and FieldML are open standards for declaratively representing mathematical models to facilitate model interchange and are primarily focussed on the needs of Physiome projects such as euHeart, 1 a cardiac modelling project with a strong focus on clinical applications.In my previous blog post, we talked about using the Color Map Editor panel to set up the color and opacity transfer functions for scalar coloring. We define a mathematical model to be a formulation that represents the state of a real-world system mathematically in such a way that the model can be used to make predictions about the real-world system by computing the state of the system based on the input parameters to the model. Functional models usually make predictions about dynamic systems. Geometric models vastly reduce the number of parameters when compared to the number of parameters required to record all the exact measurements of the real-world object. A distinction is made between implicit and explicit models. Explicit models are represented by plain numerical data and closed-form algebraic expressions, as well as certain functions commonly available in standard software math libraries, such as trigonometric functions, exponential, logarithm and so on. Implicit models include expressions that usually require the application of computational numerical solvers in order to evaluate, for example, systems of ordinary differential equations (ODEs) and partial differential equations (PDEs). The Physiome Model Repository (PMR) software provides a web repository for making models, based on these standards and other formats, easily accessible. Furthermore, PMR provides a collaboration workspace. The euHeart project aims to develop models, modelling standards and the related technologies that serve the goal of bridging the gap between cardiac modelling for research and the clinical application of individualised cardiac modelling. As part of the euHeart project, extensive design work has been done on the FieldML format and its API, and features for supporting FieldML in the PMR software. Provide open source technology to achieve the above goals. The goals of CellML have much in common, but CellML focusses on time-varying lumped parameter models. Although the format can express a wider range of models, most CellML software shares the focus on time-varying lumped parameter models. CellML focusses on the algebraic and differential mathematical equations of the model, whereas currently, FieldML focusses both on describing fields over multiple discrete indices, through reference to sparsely or densely packed 2 massive bulk numerical data, and on describing multivariate fields with some or all continuous variables, by defining finite element interpolations using the discrete data or by other interpolation methods or more general methods.Ĭomputer-readable model representation formats for Physiome models such as those used by the euHeart project need to support diverse modelling techniques. For example, typical models for cardiac mechanics simulation (e.g.
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
